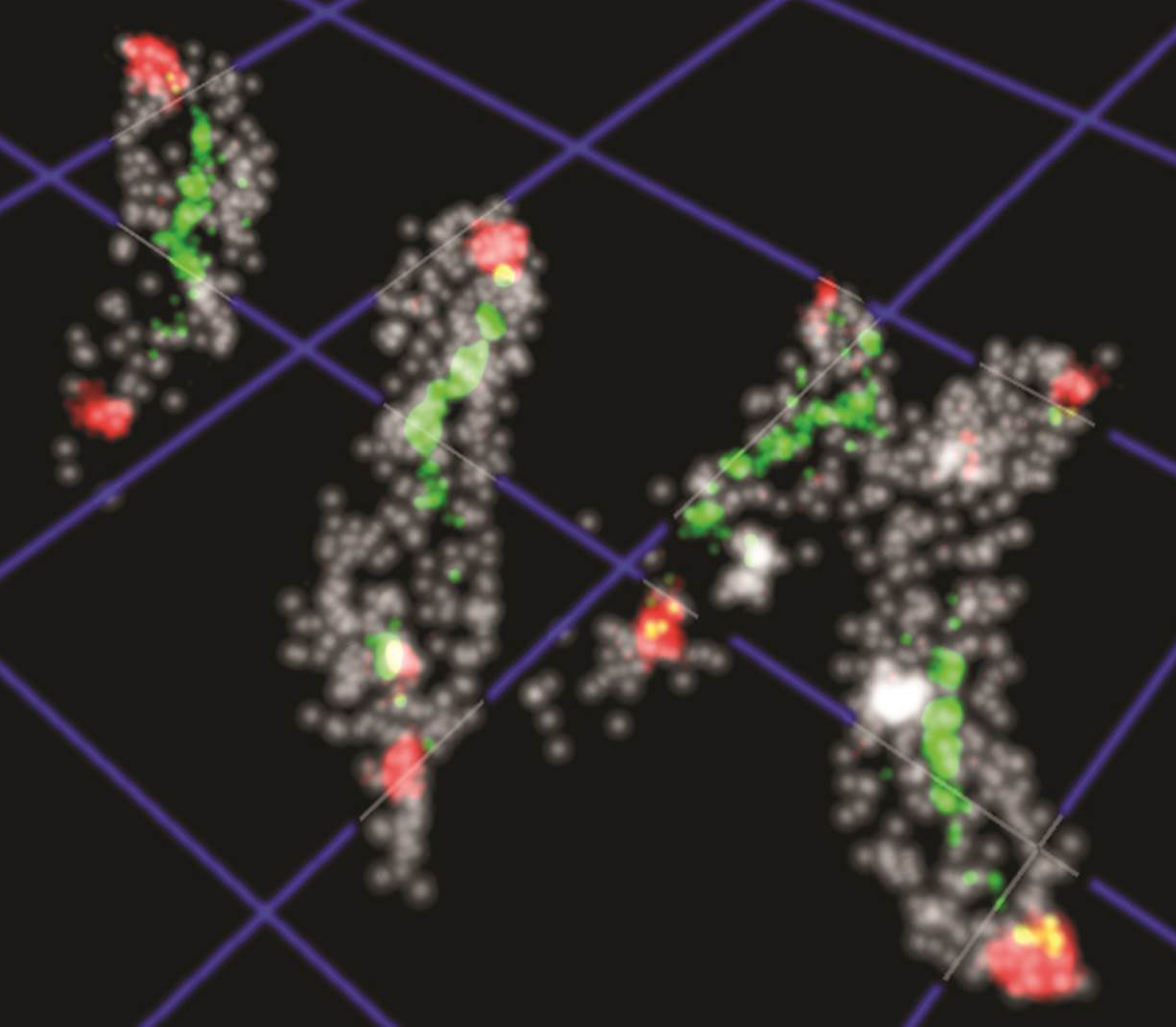
Overview
We are an interactive and multidisciplinary research lab with two primary research objectives:
Development of 3D super-resolution imaging methods, including instrument design, sample preparation, and computational image analysis.
Application of these methods to understand molecular-level spatial and temporal phenomena inside intact bacterial cells and cellular-level interactions within developing microbial communities.
To achieve these objectives, we operate in a diverse research environment that integrates aspects from several areas of chemistry, molecular and cellular biology, as well as biophysics, optical physics, engineering, and computer science. Our lab is affiliated with the Center for Membrane and Cell Physiology and the Global Infectious Disease Institute at UVA.
Cell Imaging at Relevant Length Scales
Our experiments access the molecular (nanometers) length scales that are inaccessible with conventional diffraction-limited fluorescence microscopy and bridge to the cellular (micrometers), and intercellular (10-100 micrometers) length scales. Utilizing the single-molecule sensitivity and specificity of super-resolution fluorescence microscopy, we can localize single biomolecules or single cells in 3D space and track their motion over time. Wherever possible, we primarily perform imaging experiments with living cells to characterize how molecular- or cellular-level spatial and temporal phenomena determine the physiology and phenotype of the cell or the functionalities of entire cell populations.
The molecular length scale (1 nm – 50 nm)
On this smallest length scale, we ask how specific biomolecules combine with others in their native cellular environment to produce functioning assemblies. Biomolecular assemblies often consist of several different subunits so that a set of reactions can be carried out in a spatially and temporally coordinated manner. Are the individual subunits stably or transiently incorporated? What is the temporal and spatial order of assembly? How are these biomolecular assemblies regulated at the molecular level? By measuring 3D molecular trajectories and intermolecular distances with nanometer precision, we determine how the molecular architectures of these assemblies change over time and how they are functionally regulated by the cell's developmental and signalling networks.
The cellular length scale (10 nm – 2 µm)
On the cellular length scale, we ask where biomolecules are located inside the cell. How is the population of biomolecules distributed in the cellular space? How are individual molecules moving and how is the overall distribution of the population changing in time? A large amount of evidence suggests that the appropriate positioning of many copies of a given molecule in space and time can result in emergent properties (such as cell shape maintenance or cell motility) that span length scales that are several orders of magnitude larger than the size of individual molecules. Our work determines the spatial and temporal details of the molecular mechanisms that achieve this level of control.
The intercellular length scale (200 nm – 50 µm)
On the intercellular length scale, we ask how individual cells interact with each other in a community of cells. How does a given community architecture emerge through cellular interactions or secretion of extracellular matrix material? How are social relationships among individual cells controlled through chemical or mechanical signaling between cells? While a single bacterial cell is not powerful enough to have a noticeable effect on human health, a well-organized community of bacterial cells (a biofilm) can become a thousand fold more drug resistant and substantially affect a host organism. Our research develops the super-resolution imaging and data analysis capabilities that are needed to quantitatively measure and describe how the physiology and behaviors of individual cells influence collective functions of large cell populations.
Focus on Bacterial Cell Biology
Bacteria are fundamentally interesting organisms to study at the molecular level. They are the smallest and arguably the simplest living organisms on the planet. Nonetheless, the ability to evolve rapidly in changing environments has made the bacteria an extremely diverse group of single-celled organisms. Our understanding of spatial and temporal biological phenomena inside bacteria and bacterial biofilms remains far from complete and this lack of fundamental knowledge stymies research efforts that aim to combat bacterial infections or capitalize on the vast metabolic potential found among non-pathogenic bacteria. Our imaging experiments provide a comprehensive understanding of the spatial, temporal, and chemical ecology within bacteria and bacterial biofilms, so that these organisms may be effectively utilized in biotechnological, industrial, and agricultural applications.
Bacteria are highly relevant to important economic, societal, and medicinal challenges of our time. For example, the looming inability to combat pathogenic bacteria with current antibiotics presents a major health concern. I left unresolved by the year 2050, it is anticipated that 10 Million lives will be lost to bacterial infections each year and the total disease-associated economic costs could accumulate to an estimated 100 Trillion USD from now until then (O'Neill Review on Antimicrobial Resistance). A fundamental understanding of the spatial and temporal aspects of the most widely used bacterial virulence mechanisms is therefore of utmost importance, because this knowledge will facilitate the development of new species-specific antivirulence drugs that would provide a longer lasting alternative to the broad spectrum antimicrobial drugs in use today.